DBSCAN — A Density Based Clustering Method
Do you still remember our last blog of the clustering bundle? Yes, it’s the KMeans Bundle.  In this blog, I will introduce another clustering bundle: DBSCAN Bundle — a highly scalable and parallelized implementation of DBSCAN [1] algorithm. DBSCAN is a density-based unsupervised machine learning algorithm to automatically cluster the data into subclasses or groups. Our implementation is specially designed to meet the Big Data clustering challenge by leveraging the distributed computing environment of HPCC Systems. In the following content, the essence of DBSCAN algorithm, a comparison between KMeans algorithm and DBSCAN algorithm and an example of how to use DBSCAN bundle in HPCC Systems will be introduced.
In this blog, I will introduce another clustering bundle: DBSCAN Bundle — a highly scalable and parallelized implementation of DBSCAN [1] algorithm. DBSCAN is a density-based unsupervised machine learning algorithm to automatically cluster the data into subclasses or groups. Our implementation is specially designed to meet the Big Data clustering challenge by leveraging the distributed computing environment of HPCC Systems. In the following content, the essence of DBSCAN algorithm, a comparison between KMeans algorithm and DBSCAN algorithm and an example of how to use DBSCAN bundle in HPCC Systems will be introduced.
If you are new to machine learning or interested in supervised learning, the blog Machine Learning Demystified is a great resource. It reviews the basic concepts, terminology of Machine Learning and how to utilize supervised machine learning.If you are interested in unsupervised learning and KMeans algorithm, the blog Automatically Cluster your Data with Massively Scalable K-Means is a good starter. If you are interested in the details of Myriad Interface, you may want to start from this article. This article also assumes that you have at least some basic ECL programming skills, which you can get by reading our ECL documentation or taking an online training course by clicking here.
DBSCAN Clustering Method
Density-based spatial clustering of applications with noise (DBSCAN) is a density-based clustering method. It’s well known in the machine learning and data mining communiy. DBSCAN has been widely used in both academia and industrial fields such as computer vision, recommendation systems and bio-engineering.
The principle of DBSCAN is to find the neighborhoods of data points exceeds certain density threshold. The density threshold is defined by two parameters: the radius of the neighborhood (eps) and the minimum number of neighbors/data points (minPts) within the radius of the neighborhood. With these two thresholds in mind, DBSCAN starts from a random point to find its first density neighborhood. Then from its first density neighborhood, it starts to find the second density neighborhood. If the second density neighborhood exists, DBSCAN will merge the first and second density neighborhoods to become a bigger density neighborhood. The same process keep going on until its neighborhood cannot meet the threshold anymore.
The ‘find and merge’ process allows DBSCAN to find all the density neighborhoods from a random starting point and finally merge all of them into one final big neighborhood – a cluster. After one cluster is found, DBSCAN will start the same process from another random point that has not been clustered into any group. The ‘find and merge’ process goes on until DBSCAN find another cluster and all the rest clusters. The rest data points that does not belong to any cluster are the outliers.
DBSCAN has many merits compared to other clustering methods such as KMeans. First it does not require to pre-define the number of clusters. In KMeans, the number of cluster is a fixed number as a parameter feeding into KMeans model, which means the number of clusters does not change throughout the clustering process. As we all know KMeans is an unsupervised machine learning algorithm, it’s not easy to guess the right cluster number without certain level of knowledge about the data. On the other hand, DBSCAN can automatically cluster the dataset into clusters without telling it how many clusters in the dataset. The number of clusters is changing throughout the clustering process until DBSCAN find the last cluster.
Another obvious advantage of DBSCAN is that it can detect outliers. It turns out this advantage is just a side-effect of clustering process, which does not bring any extra complex computation like some other clustering methods do. Another merits of DBSCAN is it can cluster the data in arbitrary shapes. Unlike DBSCAN, KMeans usually limits to the clusters with spherical shapes.
These characteristics make DBSCAN a good choice for anomaly detection. Because it automatically detects the number of clusters, and finds clusters of varying sizes and shapes, it can be used to identify normal usage patterns. Smaller clusters then represent infrequent patterns, and very small clusters or outliers may represent anomalies.
Now we know DBSCAN has many advantages but it also has weakness. For example, for data of varying densities, it’s hard for DBSCAN to find the right cluster and it brings challenge to set up the right model parameters. However, with all the advantages of DBSCAN, it’s worth a try to use DBSCAN in your next clustering task. In the next section, I will introduce a hands-on example of how to use DBSCAN in HPCC Systems.
Using DBSCAN Bundle in HPCC Systems
This section introduces how to use DBSCAN bundle in HPCC Systems step by step.
Step 1: Installation
- Be sure HPCC Systems Clienttools is installed on your system.
- Install HPCC Systems ML_Core Bundle
From your clienttools/bin directory run:
ecl bundle install https://github.com/hpcc-systems/ML_Core.git
3. Install the HPCC Systems DBSCAN Bundle
From your clienttools/bin directory run:
ecl bundle install https://github.com/hpcc-systems/dbscan.git
Note that for PC users, ecl bundle install must be run as Admin. Right click on the command icon and select “Run as administrator” when you start your command window.
Step 2: Import Machine Learning Bundle
Before starting running machine learning models, let’s first IMPORT them into our environment:
IMPORT DBSCAN;
IMPORT ML_Core;
Step 3: Import Raw Data
The raw data — Blobs dataset comes with the DBSCAN bundle. To import the data, simply type in below ECL code:
//Import raw data
blobs := DBSCAN.Tests.datasets.blobsDS.trainRec;
Below is a sample of the Blobs dataset:
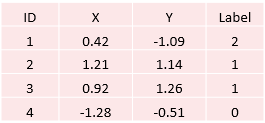
Step 4: Put Raw Data into Machine Learning Dataframe
In HPCC System, machine learning datasets are held in the dataframes defined in ML_Core bundle, such as NumericField dataframe and DiscreteField dataframe. To put our data in NumericField dataframe, you can type in below source code:
// Convert raw data to NumericField format
ML_Core.ToField(blobs,recs);
Step 5: Run DBSCAN model
As introduced previously, DBSCAN model has a few parameters such as ‘esp’ and ‘minPts’. Here we set the parameter esp = 0.3 and the minPts = 2 to define the model as blew. However this step is not required. It’s totally fine to ignore them and use the default values. You can find details about other optional parameter such as ‘distance_func’ in the documentations.
//Train DBSCAN model
Model := DBSCAN.DBSCAN(0.3, 2).Fit( recs(number < 3));
Note: Here we filter out the unnecessary attributes out via filter (number < 3).
After the model is built, we can take a look how many clusters in the dataset:
NumClusters := DBSCAN.DBSCAN().Num_Clusters(Model);
One important feature of DBSCAN is to detect outliers in our dataset. Let’s take a look how many outliers in the Blobs dataset and who they are.
NumOutliers := DBSCAN.DBSCAN().Num_Outliers(Model);
Outliers := Model(label = 0);
Conclusion
Congrats you just learned another high performance and easy-to-use clustering bundle DBSCAN on the HPCC Systems platform! In this article, I introduced the principle of how DBSCAN works, the Pros and Cons of its clustering method and a hands-on tutorial. Hope our DBSCAN bundle and this article is helpful for your next Machine Learning project.
Find out more…
- Read Roger Dev’s introductory blog which focuses on Demystifying Machine Learning.
- Find out about Using PBblas on HPCC Systems
- We have restructured the HPCC Systems Machine Learning Library. Find out more about this and future improvements.
- Contribute to our Machine Learning library and join our active HPCC Systems community
Reference
[1] Ester, Martin, et al. “Density-based spatial clustering of applications with noise.” Int. Conf. Knowledge Discovery and Data Mining. Vol. 240. 1996.
